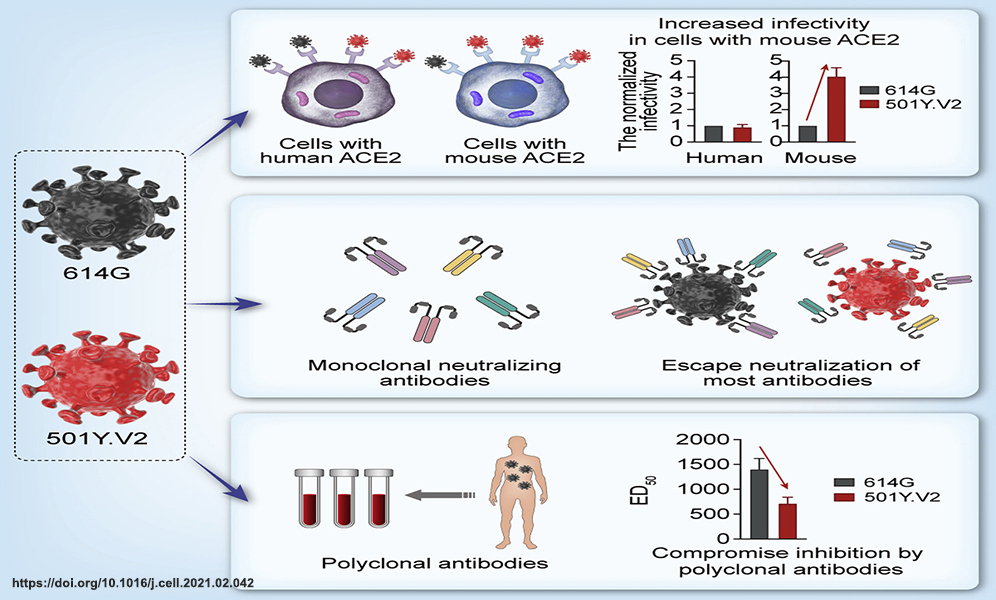
The mutation is a common phenomenon in the natural evolution of viruses, and SARS-CoV-2 is no exception. In contrast to the rapid evolution of other RNA viruses, SARS-CoV-2 has low genomic variability because of its proofreading function. Therefore, to try to control the pandemic, researchers rushed to make vaccines and monoclonal antibodies (mAbs) tailored to the viral sequence initially identified in Wuhan, China. Thanks to incredible technological progress, huge public-sector investments, and global scientific collaboration, vaccines with the efficacy of up to 95% and lifesaving mAbs were developed and approved for emergency use at a speed never achieved before. However, this ray of hope was clouded by the emergence of a variety of SARS-CoV-2 mutants during the ongoing pandemic. Mutants may be more transmittable or may be able to evade neutralizing antibodies induced by either infection or vaccination, so-called immune escape.
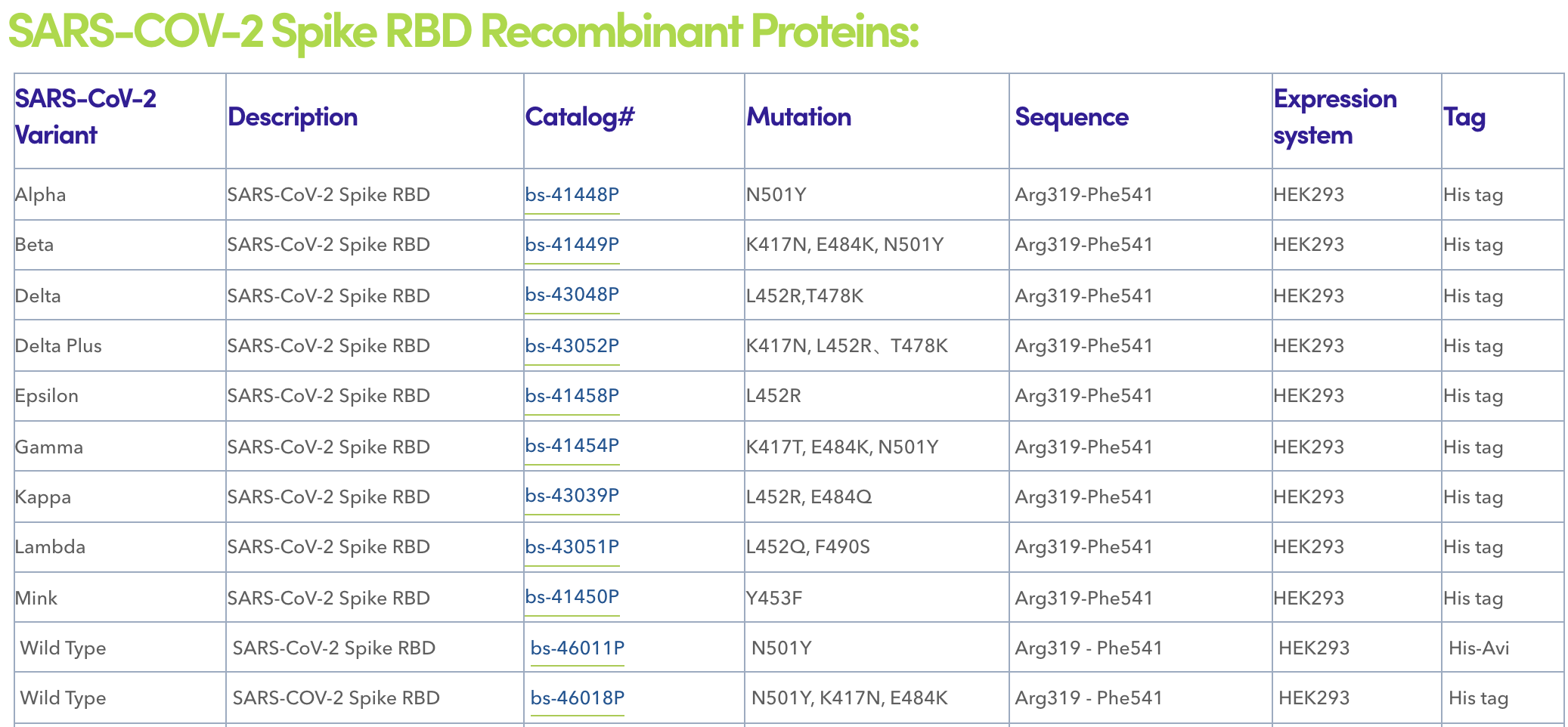
Serological analyses of almost 650 individuals infected with SARS-CoV-2 indicated that ~90% of the plasma or serum neutralizing antibody activity targets the spike receptor-binding domain (RBD). A relative lack of glycan shielding may contribute to the immunodominance of the RBD. Structural analyses allowed the categorization of RBD-binding neutralizing antibodies into four classes: ACE2-blocking antibodies that bind the spike protein in the open conformation (class 1); ACE2-blocking antibodies that bind the RBD in both the open conformation and the closed conformation (class 2); antibodies that do not block ACE2 and bind the RBD in both the open conformation and the closed conformation (class 3); and neutralizing antibodies that bind outside the ACE2 site and only in the open conformation (class 4). Interestingly, further studies on how mutations in the SARS-CoV-2 spike protein affect antibodies neutralization found that the mutations that reduce antibody binding the most occur at a relatively small number of RBD residues. These findings together indicate the critical role of RBD in immune defense and immune escape. Nowadays, multiple RBD mutations have been identified in the clinic. Among them, E484 has been considered as a key escape mutation target, with amino acid changes to K, Q, or P, reducing antibodies neutralization. It was reported that some other mutations at position K417, L452, and N501 may also cause alleviated neutralizing antibody binding and immune escape.
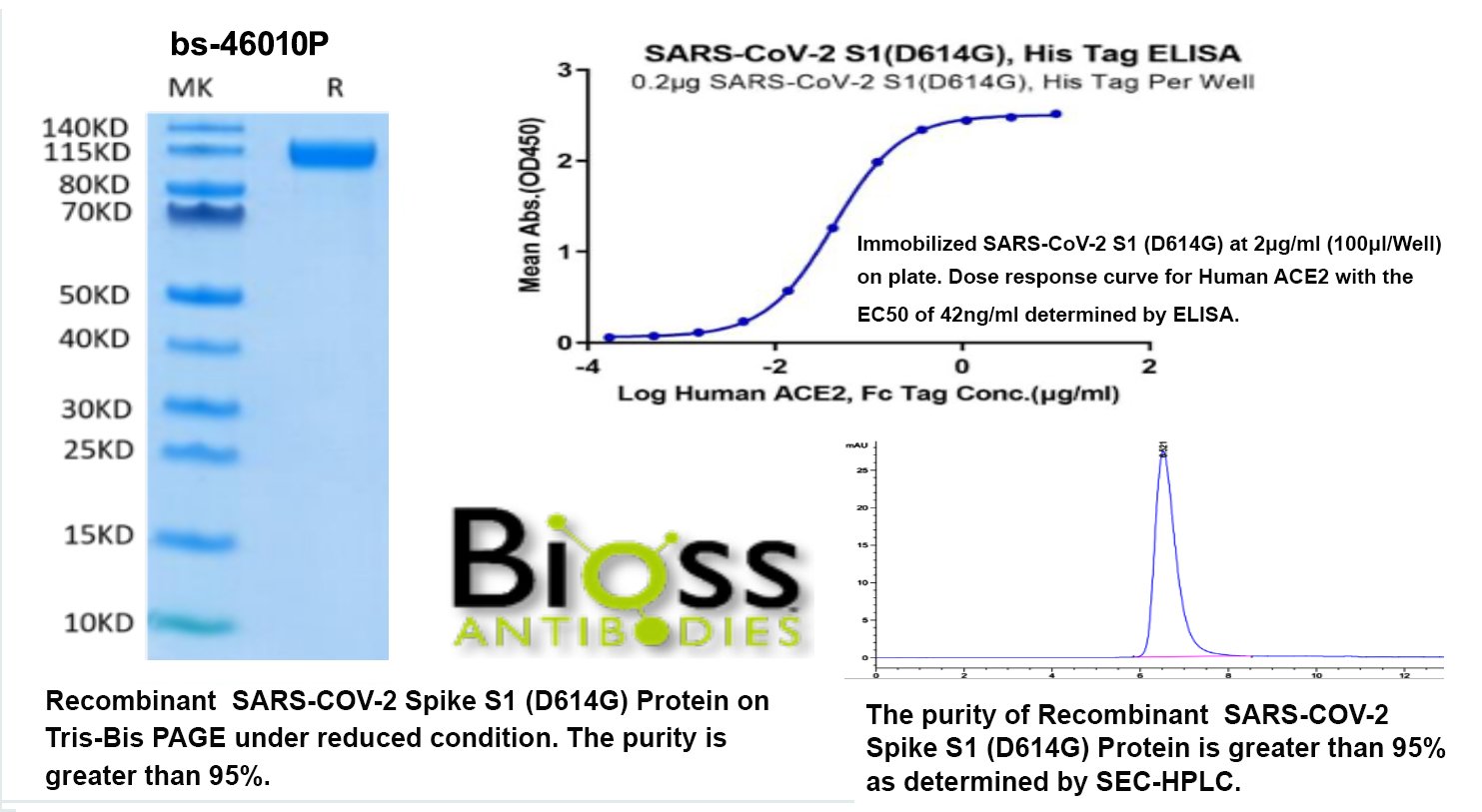
It is clear that RBD amino acid substitutions impact neutralizing antibodies in the global virus population, and there is emerging evidence of RBD-mutation-associated variants exhibiting resistance to antibody-mediated immunity elicited by vaccines. To control the pandemic, a greater understanding of the correlates of immune protection is required to provide a context for the results of studies reporting changes in neutralization. A comprehensive understanding of the consequences of spike mutations, especially RBD mutations for antigenicity, will encompass both T cell-mediated immunity and non-spike epitopes recognized by antibodies. More generally, a broader understanding of the phenotypic impacts of mutations across the SARS-CoV-2 genome and their consequences for variant fitness will help elucidate drivers of transmission and evolutionary success. To this end, many organizations, including Bioss, are dedicated to developing and manufacturing various SARS-CoV-2 viral proteins.
----------------------------------------------------------
References:
1. Harvey, W.T., Carabelli, A.M., Jackson, B. et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol 19, 409–424 (2021). https://doi.org/10.1038/s41579-021-00573-0
2. Li, Q., Nie, J., Wu, J. et al. SARS-CoV-2 501Y.V2 variants lack higher infectivity but do have immune escape. Cell 184, 2362-2371.e9 (2021). https://doi.org/10.1016/j.cell.2021.02.042.